Archive
Prediction problem
In class, I described a simple Bayesian prediction problem. One observes the number of fire calls in a small California city for 30 days — the total number of fire calls in this period is 158. An administrator is interested in predicting the total number of calls in the next seven days.
We assume that y, the number of calls in a day, is Poisson with mean
. Suppose our prior beliefs about
are described by a normal density with mean 10 and standard deviation 2.
I first write a simple function compute.post that computes and plots the posterior (using the likelihood times prior recipe) on a grid between lo and hi.
compute.post=function(lo, hi,...){
lambda = seq(lo, hi, length=100)
prior = dnorm(lambda, 10, 2)
like = lambda^158 * exp(-30*lambda)
post = prior * like
plot(lambda, post,...)
list(L = lambda, P = post)
}
With some trial and error, the interval (3.3, 7.5) seems to bracket the posterior well.
S=compute.post(3.3, 7.5, type="l", col="red")
Next I take a simulated sample from the posterior by using the sample function.
sim.lambda = sample(S$L, 1000, replace=TRUE, prob=S$P)
Last, I simulate a sample from the posterior predictive density of z, the number of fire calls in seven days. Since [z | ] is Poisson(7
, we simulate z by (1) simulating
from the posterior and (2) simulating z from a Poisson(7
) distribution. This is easy to do by using the vector properties of the random simulation functions.
sim.z = rpois(1000, 7*sim.lambda)
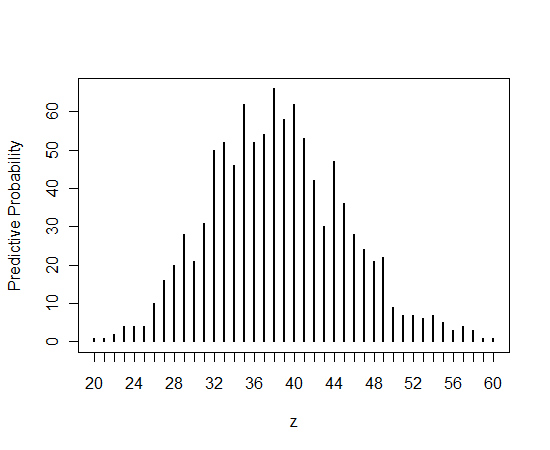
Here is a graph of the predictive distribution (by tabulating and plotting the simulated values of z).
Brute force Bayes for one parameter
Although we talk a lot about conjugate analyses, one doesn’t need to restrict oneself to the use of conjugate priors. Here we illustrate learning about a Poisson mean using a normal prior on the mean .
Suppose I observe the following number of fire calls for a particular community each week: 0, 0, 1, 3, 2, 2. If we assume , the number of fire calls in week
, is Poisson(
), then the likelihood function is given by
Suppose I represent my prior beliefs about by a normal curve with mean 3 and standard deviation 1. Then the posterior is given by (up to an unknown proportionality constant) by
Here is a simple brute-force method of summarizing this posterior.
1. Choose a grid of values that covers the region where the posterior is concentrated (this might take some trial and error).
2. On this grid, compute the prior, likelihood, and posterior.
3. Using the R sample function, take a large sample from the grid where the probabilities of the points are proportional to the like x prior values.
4. Summarize this posterior simulated sample to learn about the location of the posterior.
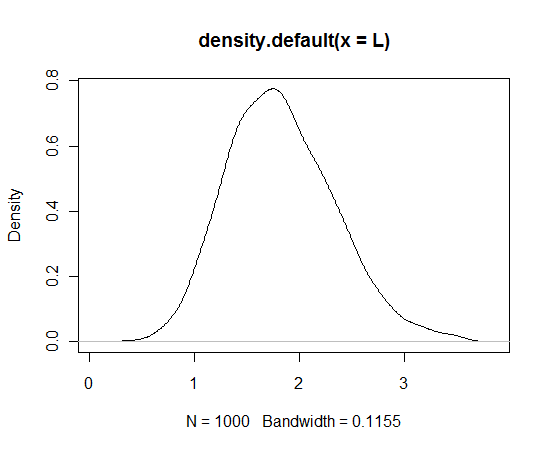
Here’s some R code for this example. I use the plot function to make sure the grid does cover the posterior. The vector L contains 1000 draws from the posterior.
lambda = seq(0, 5, by=0.1) like = exp(-6*lambda)*lambda^8 prior = dnorm(lambda, 3, 1) post = like * prior plot(lambda, post) L = sample(lambda, size=1000, prob=post, replace=TRUE) plot(density(L)) # density graph of simulated draws
Poisson mean example, part II
In the last post, I described the process of determining a prior for , my son’s average number of text messages per day. I decided to model my beliefs with a gamma(a, b) density with a = 44.4 and b = 4.4.
Now I observe some data. I looked at the online record of text messaging for the first seven days that’s been at school and observe the counts
Sat Sun Mon Tue Wed Thu Fri
19 4 26 17 15 0 17
If we assume these counts are Poisson with mean , then the likelihood function is given by
L(lambda) = lambda^s exp(-n lambda),
where is the number of observations (7) and
is the sum of the observations ( 98).
The posterior density (using the prior times likelihood recipe) is given by
L(lambda) x g(lambda) = lambda^{a+s-1} exp(-(b+n) lambda),
which we recognize as a gamma density with shape and rate
.
Using the following R commands, we construct a triplot (prior, likelihood, and posterior).
> a=44.4; b=4.4
> s=sum(y); n=length(y)
> curve(dgamma(x,shape=s+a,rate=n+b),col=”red”,xlab=”LAMBDA”,
+ ylab=”DENSITY”,lwd=3,from=3,to=25)
> curve(dgamma(x,shape=a,rate=b),col=”blue”,lwd=3,add=TRUE)
> curve(dgamma(x,shape=s+1,rate=n),col=”green”,lwd=3,add=TRUE)
> legend(“topright”,c(“PRIOR”,”LIKELIHOOD”,”POSTERIOR”),
+ col=c(“blue”,”green”,”red”),lty=1,lwd=3)
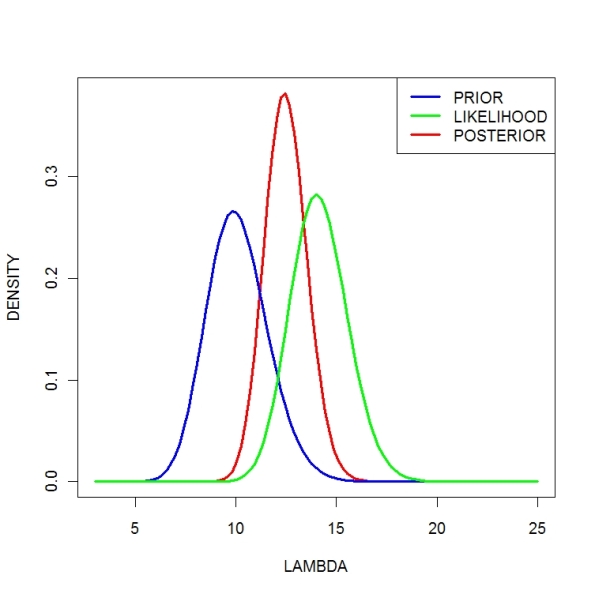 Note that (1) the posterior is a compromise between the data information (likelihood) and the prior and (2) the posterior is more precise since we are combining two sources of information.
Note that (1) the posterior is a compromise between the data information (likelihood) and the prior and (2) the posterior is more precise since we are combining two sources of information.> ys=rpois(1000,30*lambda)
> plot(table(ys),ylab=”FREQUENCY”)
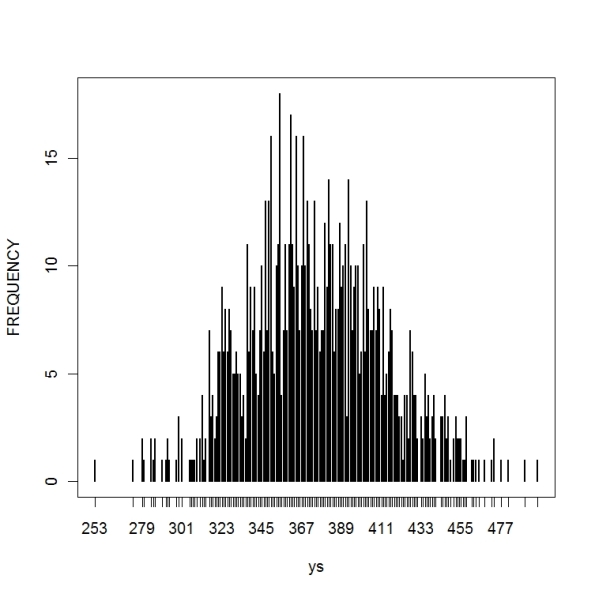 Looking at the graph, it is likely that my son will have between 310 and 450 text messages in the next 30 days.
Looking at the graph, it is likely that my son will have between 310 and 450 text messages in the next 30 days.Constructing a prior for a Poisson mean
I am interested in learning about my son’s cell phone use. Suppose the mean number of text messages that he sends and receives per day is equal to . I will observe the number of text messages
for
days — we’ll assume that conditional on
,
are independent Poisson(
).
How do I construct a prior density on ? Suppose I model my beliefs by use of a gamma density with shape parameter
and rate parameter
. I want to figure out the values of the prior parameters
and
.
There are two distinct ways of assessing your prior. One can think about plausible values of the mean number of daily text messages . Alternatively, it may be easier to think about the actual number of text messages
— if we assume a gamma prior, then one can show that the predictive density of
is given by
Personally, I think it is easier to think about the number of text messages . My best prediction at
is 10. After some additional thought, I decide that my standard deviation for
is equal to 3.5. One can show that the mean and standard deviation of the predictive density are given by
If one matches my guesses with these expressions, one can show .
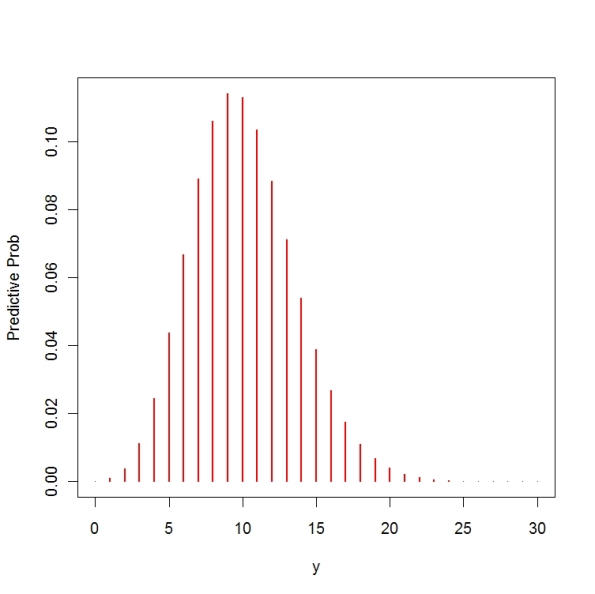
To see if these values make any sense, I plot my predictive density. This density is a special case of a negative binomial density where (using the R notation)
size = a, prob = b /(b + 1).
I graph using the following R commands. One can compute P(5 <= y <= 15) = 0.89 which means that on 89% of the days, I believe Steven will send between 5 and 15 messages.
a = 44.4; b = 4.4
plot(0:30,dnbinom(0:30,size=a,prob=b/(b+1)),type=”h”,
xlab=”y”, ylab=”Predictive Prob”, col=”red”, lwd=2)
Modeling with Cauchy errors
One attractive feature of Bayesian thinking is that one can consider alternative specifications for the sampling density and prior and these alternatives makes one think more carefully about the typical modeling assumptions.
Generally we observe a sample distributed from a sampling density
depending on a parameter
. If we assign
a prior density
, then the posterior density is given (up to a proportionality constant) by
Consider the following Cauchy/Cauchy model as an alternative to the usual Normal/Normal model. We let be distributed from a Cauchy density with location
and known scale parameter
, and we assign
a Cauchy prior with location
and scale
.
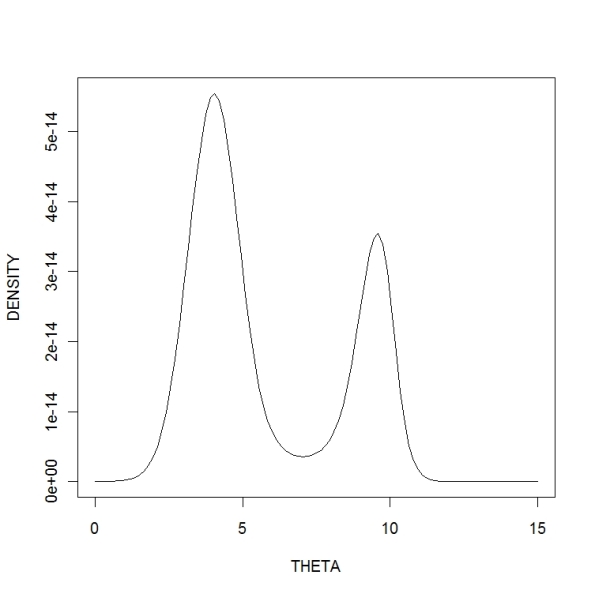
Of course, this is not going to be a conjugate analysis — the posterior density has a complicated form. But it is easy to use R to plot and summarize the posterior density.
First we define the posterior density for . The arguments to the function are theta (which could be a vector), the vector of data values, the known scale parameter
, and the prior parameters
andn
. The output is a vector of values of the (unnormalized) posterior density.
posterior=function(theta,data,scale,mu0,tau)
{
f=function(theta) prod(dcauchy(data,theta,scale))
likelihood=sapply(theta,f)
prior=dcauchy(theta, mu0, tau)
return(prior*likelihood)
}
How many more home runs will Ryan Howard hit?
Here is a basic proportion problem. We are approaching the end of the 2009 baseball season and Ryan Howard of the Philadelphia Phillies currently has 38 home runs in 540 at-bats (opportunities). I project that he will have 85 more at-bats this season. How many additional home runs will he hit?
Let p be the probability that Howard hits a home run in a single at-bat during the 2009 season. We’ll address this prediction problem in three steps. First, we’ll use past data from previous seasons to form a prior distribution for p. Second, we’ll update our beliefs about p using the 2009 data. Last, we’ll predict the number of future at-bats from the posterior predictive distribution. In this process, we’ll illustrate the use of some functions from the LearnBayes package in R.
1. My prior.
In the four previous seasons (2005 through 2008) his home runs/at-bats have been 22/312. 58/581. 47/529, and 48/610. The home run rates are 0.071, 0.100, 0.089, 0.079. Remember that p is the 2009 home run hitting probability. Based on these data, I believe that
(1) P(p < 0.075) = 0.5
(2) P(p < 0.115) = 0.9
In other words, my best guess at p is 0.075 and I’m pretty confident that p is smaller than 0.115. I find the beta(a, b) that matches this information by use of the beta.select function in LearnBayes. I first define the two quantiles as lists — these quantiles are the arguments in beta.select.
> quantile1=list(p=.5,x=.075)
> quantile2=list(p=.9,x=.115)
> beta.select(quantile1, quantile2)
[1] 7.26 85.77
> n=85
> y=0:85
> pred.probs=pbetap(ab,n,y)
Do graduate students prefer McDonalds?
Today I did an illustration of discrete Bayes for a proportion. I’m interested in the proportion p of graduate students who answer “McDonalds” when asked the question “McDonalds, Wendys, or Burger King?”
I believe p can be one of the five values 0.1, 0.2, 0.3, 0.4, 0.5 and I assign the respective prior weights 1, 2, 5, 10, 5. I define this prior in R:
> p=c(.1,.2,.3,.4,.5)
> prior=c(1,2,5,10,5)
> prior=prior/sum(prior)
> names(prior)=p
> barplot(prior,ylim=c(0,.6),xlab=”p”,main=”PRIOR”)
> plot(s,xlab=”p”,main=”POSTERIOR”)
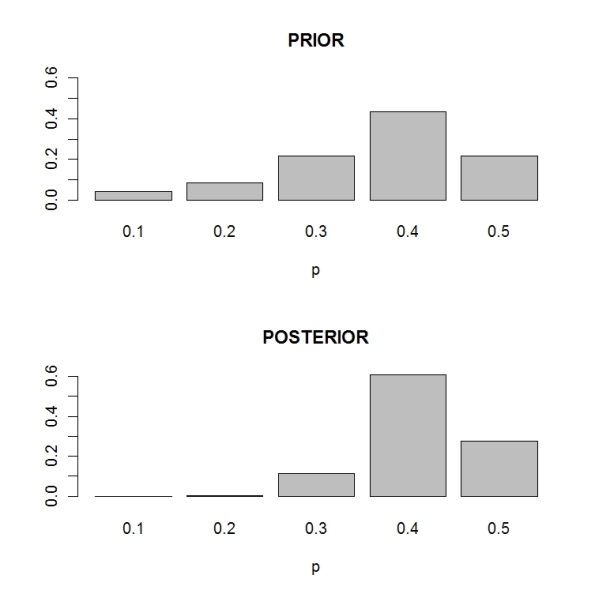 Note that the posterior probs are more precise than the prior probabilities. I am more confident that the proportion of McDonalds fans is equal to 0.4.
Note that the posterior probs are more precise than the prior probabilities. I am more confident that the proportion of McDonalds fans is equal to 0.4.Discrete Bayes, part II
In the previous post, I used the R function dexp in my discrete Bayes function. Sometimes you need to make slight changes to the functions in the R stats package for a given problem.
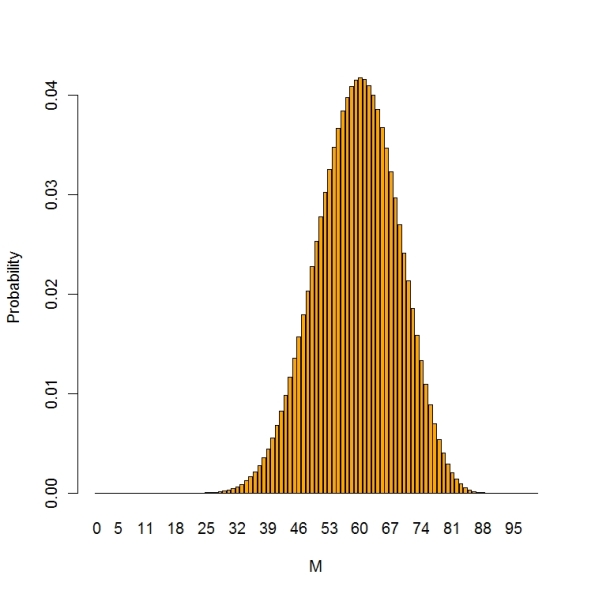
Here is a finite population sampling problem. A small community has 100 voters and you are interested in learning about the number, say M, who will vote for a school levy. You take a random sample of size 20 (without replacement) and 12 are in support of the levy. What have you learned about M?
Here the sampling density is hypergeometric — the probability that will be in favor out of 20 given that M in the population favor the levy — this is given by the hypergeometric probability
There is a function dhyper that computes this probability, but uses a different parametrization. So I write a new function dhyper2 that uses my parametrization.
dhyper2=function(y,M,sample.size=n,pop.size=N)
dhyper(y,M,N-M,n)
Discrete Bayes
One good way of introducing Bayesian inference is by the use of discrete priors. I recently wrote a simple generic R function that does discrete Bayes for arbitrary choice of a prior and sampling density. I’ll illustrate this function here and in the next posting.
Suppose I’m interested in learning about the rate parameter of an exponential density. We observe a sample from the density defined by
where we are interested in learning about the rate parameter .
In R, I read in the function “discrete.bayes” and a few associated methods by typing
> source(“http://bayes.bgsu.edu/m6480/R/discrete.bayes.functions.R”)
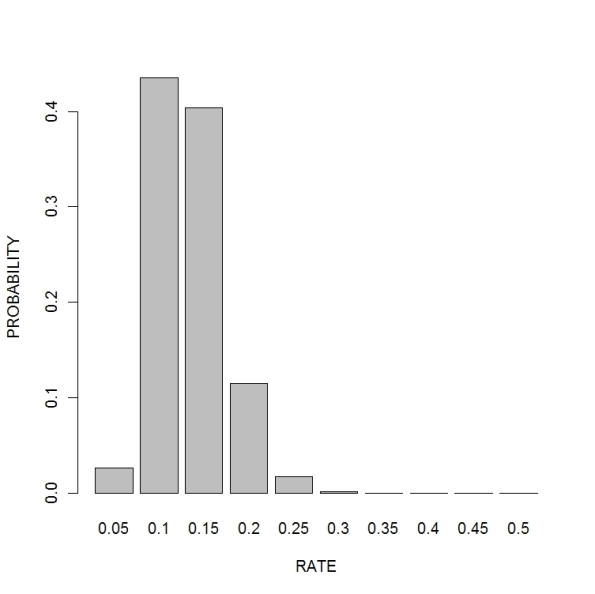 To find a 90% probability estimate for the rate, use summary:
To find a 90% probability estimate for the rate, use summary:Sequential learning by Bayes’ rule
I’m talking about Bayes’ rule now and here’s a quality control example (I think it comes from an intro Bayesian book by Schmitt that was published in the 1960’s):
A machine in a small factory is producing a particular automotive component. Most of the time (specifically, 90% from historical records), the machine is working well and produces 95% good parts. Some days, the machine doesn’t work as well (it’s broken) and produces only 70% good parts. A worker inspects the first dozen parts produced by this machine on a particular morning and obtains the following results (g represents a good component and b a bad component): g, b, g, g, g, g, g, g, g, b, g, b. The worker is interested in assessing the probability the machine is working well.
Here we have two models: W, the machine is working, and B, the machine is broken, and we initially believe P(W) = 0.9 and P(B) = 0.1. We know the characteristics of each machine. If the machine is working, it produces 95% good parts, and if the machine is broken, it produces only 70% good parts.
We use the odds form of Bayes’ rule:
Posterior Odds of W = (Prior Odds of W) x BF,
where BF (Bayes factor) is the ratio P(data | W)/P(data|B). It is convenient to express this on a log scale:
log Post Odds of W = log Prior Odds of W + log BF
Let’s illustrate how this works. Initially the prior odds of working is
log Prior Odds = log [P(W)/P(B)] = log(0.9/0.1) = 2.20.
We first observe a good part (g). The log BF is
log BF = log (P(g | W)/log(g | B)) = log(0.95/0.70) = 0.31,
so the new log odds of working is
log Posterior Odds = 2.20 + 0.31 = 2.51
so we are more confident that the machine is in good condition.
We can continue doing this in a sequential fashion. Our posterior odds becomes our new prior odds and we update this odds with each observation. If we observe a good part (g), we’ll add 0.31 to the log odds; if we observe a bad part (b), we’ll add
log [P(b | W) / P(b | B)] = log(0.05/0.30) = -1.79
to the log odds. I did this for all 12 observations. Here’s a graph of the results (first I’ll show the R code I used).
d=c(“g”,”b”,”g”,”g”,”g”,”g”,”g”,”g”,”g”,”b”,”g”,”b”)
log.BF=ifelse(d==”g”,log(.95/.70),log(.05/.30))
prior.log.odds=log(.9/.1)
plot(0:12,c(prior.log.odds,prior.log.odds+cumsum(log.BF)),
type=”l”,lwd=3,col=”red”,xlab=”OBSERVATION”,
ylab=”log odds of working”,main=”Sequential Learning by Bayes’ Rule”)
abline(h=0)
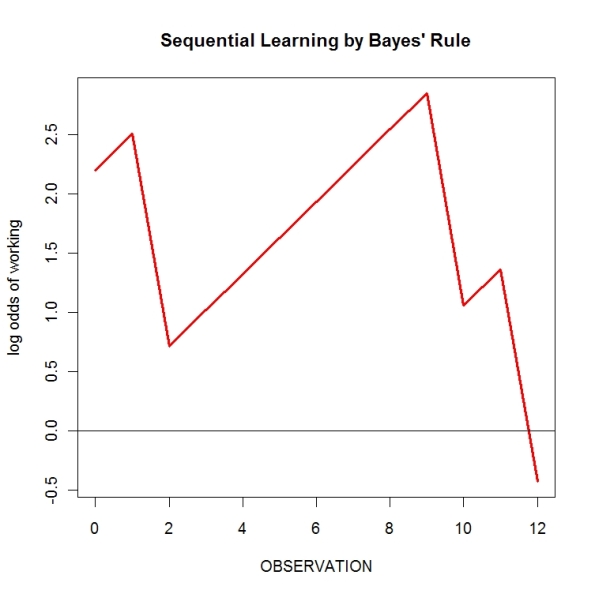 After observation, the posterior log odds is -0.43 which translates to a posterior probability of exp(-0.43)/(1+exp(-0.43)) = 0.39 that the machine is working. The machine should be fixed.
After observation, the posterior log odds is -0.43 which translates to a posterior probability of exp(-0.43)/(1+exp(-0.43)) = 0.39 that the machine is working. The machine should be fixed.